Our research
Our research group is interested in the molecular mechanisms defining the transcriptomic and epigenomic states of oligodendrocyte lineage cells. We are particularly focused on how interplay between transcription factors and chromatin modulators contributes to the transition between epigenetic states in oligodendroglia, with the aim to design epigenetic based-therapies to induce regeneration/remyelination and prevent neuroinflammation in demyelinating diseases, such as MS.
Research area
All cells in a given organism are derived from a single cell (zygote) and thereby share an identical genome. Additional layers of epigenetic information overlaid on the genome achieve the plethora of cellular phenotypes present in development and in the adult body. This epigenetic information is stored at the level of chromatin, the complex where nuclear DNA is packaged together with histones. DNA methylation and post-translational modifications at histones define the epigenetic state of a cell and ultimately cell fate, by controlling key processes, including transcription.
Oligodendrocytes insulate neuronal axons through their myelin containing membranes. Myelin allows the fast and efficient impulse transmission between neurons through saltatory conduction and is important for axonal integrity, thereby being essential for the proper functioning of the central nervous system.
Several diseases, such as multiple sclerosis (MS), are characterized by abnormal or defective myelination. Spontaneous remyelination occurs at initial stages of MS, promoted by endogenous oligodendrocyte precursor cells (OPCs). However, this process progressively starts occurring with less efficiency, until it eventually fails. Oligodendrocyte precursors (OPCs) start to be specified early during embryogenesis, in different areas of the embryonic brain, but their terminal differentiation and functional maturation occurs only at post-natal stages. The epigenetic state of OPCs define their ability to remain as a precursor cell, differentiate or even de-differentiate into a stem cell state or a glioma initiating cell state.
The main focus of our research group is to investigate how different epigenetic states in oligodendroglia (OPCs and oligodendrocytes) are established, by identifying key chromatin modulators that are involved in epigenetic transitions, using technologies such as single-cell and spatial transcriptomics and epigenomics, among others. We identified novel distinct mouse and human oligodendrocyte cell states during differentiation/myelination and also during development (Science 2015, Science 2016, Developmental Cell, 2018, Nature Communications 2020, Developmental Cell 2022, Nature Neuroscience 2026). We have also expanded our single-cell omics approach to the MOG EAE mouse model of MS and to human patient samples, in which we could discriminate distinct disease-associated OPC/OL states (Nature Medicine, 2018, Nature 2019, Neuron 2024). We have implemented scATAC-Seq, which allows examination of chromatin accessibility at a single cell level, applying for instance to examine the oligodendrocyte lineage in the EAE mouse model of MS (Neuron 2022). We found that oligodendroglia presents epigenomic priming at immune genes, to allow their transcription in inflammatory environments, but might also present epigenetic memory at later stages (Neuron 2022, Nature Neuroscience 2025). We also found that some susceptibility variant for MS are located in regions of open chromatin in human and mouse oligodendroglia, which might implicates these cells in MS aetiology and progression.
In collaboration with Prof. Mats Nilsson at Stockholm University, we have applied in situ sequencing to unveil disease evolution in the EAE mouse model of MS and unveil new neuropathological compartments within lesion on archival spinal cord tissue from MS patients (Cell 2024). We are very interested in technological development in single-cell and spatial epigenomics. We have developed single-cell CUT&Tag to examine the simultaneous profile of histone modifications at a single-cell level in large number of cells and applied to investigate the epigenomic heterogeneity of the mouse brain (Nature Biotechnology 2021). We also developed a multimodal and optimized iteration of scCUT&Tag called nano-CT (for nano-CUT&Tag) that allows simultaneous probing of three epigenomic modalities at single-cell resolution, using nanobody-Tn5 fusion proteins (Nature Biotechnology 2023). In collaboration with Prof. Rong Fan at Yale University, we have implemented CUT&Tag and ATAC-Seq at a spatial level, by using a new ligation-based method for deterministic barcoding in tissue, probing different histone modifications and chromatin accessibility in the mouse brain (Nature 2022 and Science 2022). Together we have also implemented spatial multiomics approaches for RNA/epigenomics/protein (Nature 2023, Nature 2025).
We generated several web resources from our single-cell and spatial omic datasets, compiled in our OligoInternode interface, where one can enter your gene of interest and investigate its expression pattern in the identified populations and cell states, including the oligodendrocyte lineage. One can explore how a gene of interest is differentially expressed or the chromatin landscape in regulatory regions surrounding your gene of interest. We have also developed KaroSpace (link https://karospace.se/), with which you can quickly generate a browser to explore your own or published spatial omics data.
Our lab is also part of KNIMS - The Karolinska Neuroimmunology & Multiple Sclerosis Network
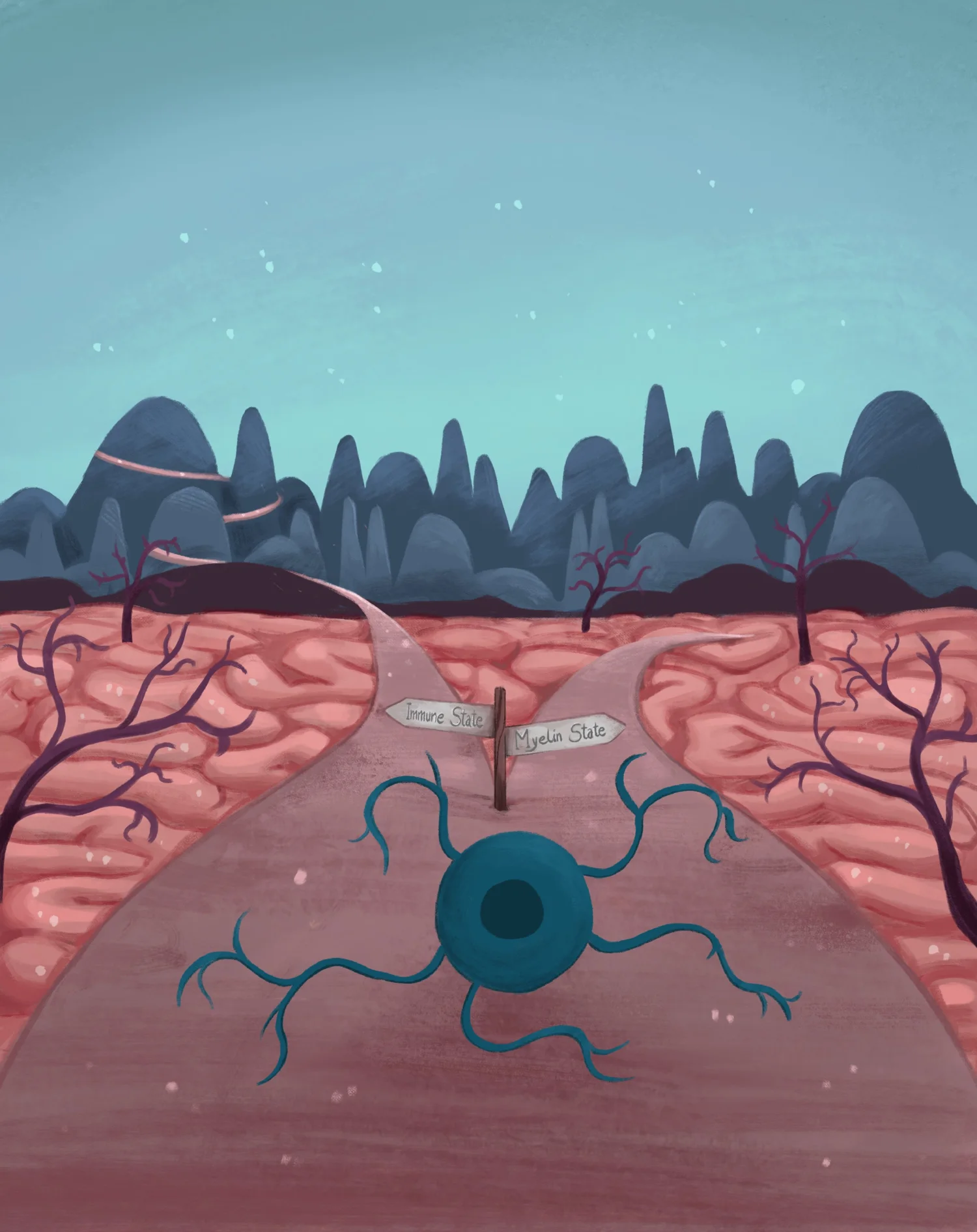

 Photo: SciLifeLab
Photo: SciLifeLabOur research group is a part of SciLifeLab
SciLifeLab is a joint effort involving multiple Swedish universities that takes part in collaborations with healthcare, industry, governmental agencies, and international organizations. This state-of-the-art national research infrastructure and research environment for molecular biosciences provides advanced technologies and expertise for fundamental and applied research in the life sciences.