Our research
Neurodegenerative diseases are characterized by the selective loss of specific neuronal populations with corresponding distinct clinical features. Why specific neuron types are selectively vulnerable to a neurodegenerative disease process, such as motor neurons in amyotrophic lateral sclerosis (ALS) or dopamine neurons in Parkinson’s disease (PD), is currently unclear. It is particularly intriguing as disease-causative genes are often broadly expressed in the nervous system and sometimes even in every cell in our body.
We are focused on understanding mechanisms of selective neuronal vulnerability and resilience seen in neurodegenerative diseases, with particular emphasis on the lethal motor neuron disease ALS. We believe that investigating cell intrinsic properties of neurons that show either extreme vulnerability or particular resilience and even regenerative properties in response to disease could reveal mechanisms that can be targeted by gene therapy approaches to render neurons more resistant.
Towards this purpose we have developed a method (LCM-seq) that allows for spatial RNA sequencing at the single cell level of partially degraded human tissues with exceptional sensitivity and gene detection. We are now using LCM-seq to unravel motor neuron diversity in the human spinal cord as well as to deduce the response of individual neuron to disease as these are either degenerating, persisting or regenerating in response to ALS.
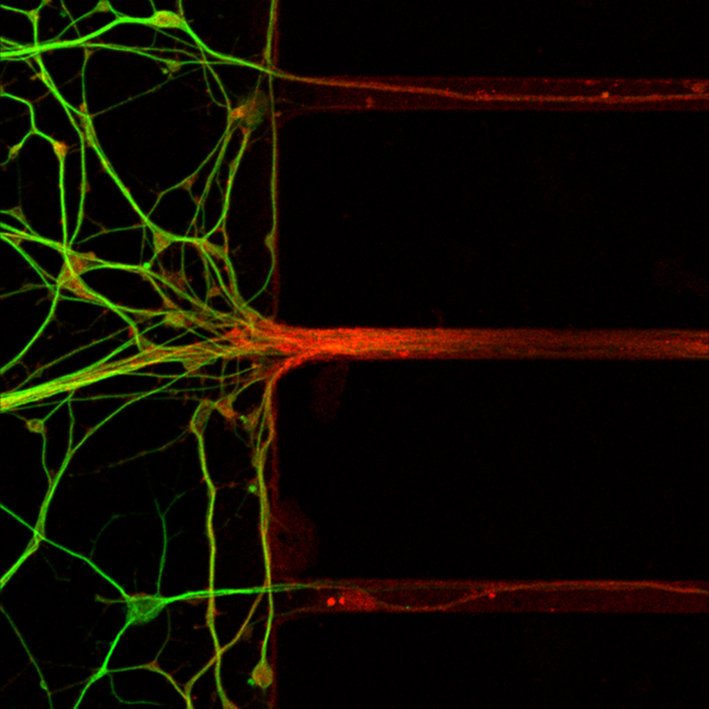
In several neurodegenerative diseases, it is the long neuronal processes (axons) and their synapses (with other neurons or muscle) that first show signs of pathology and degenerate. Still, these axonal processes are often overlooked. Towards the goal of understanding early disease processes in axons and identify targets for disease intervention, we have developed a highly robust method (Axon-seq) to sequence the content of axons and to analyze disease-induced dysregulation of the axonal mRNA content
We are now utilizing Axon-seq to increase the understanding of axon biology in general and to unravel early disease mechanisms in ALS using neurons specified from human induced pluripotent stem cells where we have introduced disease-causing mutations using CRISPR-Cas9 genome editing.
News archive
More news in Swedish
Hedlund lab on twitter
Follow us...
