Protein misfolding and prevention in cancer
Gastrokine 1 (GKN1) and GKN2 are both expressed in the gastric epithelium, and GKN1 and GKN2 levels are decreased significantly in gastric cancer. GKN1 and GKN2 both contain the chaperone-like domain BRICHOS. Our research line will focus on functions and mechanisms of this domain for potential novel approaches against cancer.
The tumor suppressor protein p53 plays an important role in suppressing cancer, while defective p53 is linked to cancer development. As many as 60% of all human tumors (including gastric cancer) contain mutated p53, and consequently, restoring its function would be a major step in cancer therapy. This project will study whether BRICHOS from GKN1 and GKN2 can restore the activity of abnormal p53.
Due to its partially disordered nature, so far, no high-resolution structure of the full-length p53 has been determined. This project, in collaboration with the groups of Michael Landreh, Philip Köck aims to solve the p53 structure at atomic resolution by cryogenic electron microscopy (cryo-EM).
Protein aggregation and prevention in Alzheimer’s disease
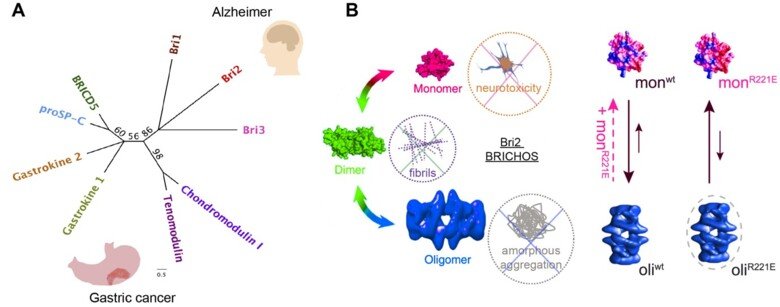
The two core pathological hallmarks of Alzheimer’s disease (AD) are intracellular neurofibrillary tangles and amyloid plaques, resulting from fibrillar-aggregation of tau and amyloid-β peptide (Aβ), respectively. The key challenge in finding an effective treatment is to specifically interfere with tau and/or Aβ fibrillar-aggregation and subsequently alleviate neurotoxicity. Understanding the molecular mechanisms behind these phenomena may have important impact on the possibilities to treat also other protein misfolding diseases. This project will focus on molecular mechanisms of Bri2 BRICHOS against self-assembly of proteins and peptides, in particular tau.
Augmentation of Bri2 molecular chaperone activity against amyloid-β reduces neurotoxicity in mouse hippocampus in vitro.
Chen G, Andrade-Talavera Y, Tambaro S, Leppert A, Nilsson HE, Zhong X, Landreh M, Nilsson P, Hebert H, Biverstål H, Fisahn A, Abelein A, Johansson J
Commun Biol 2020 01;3(1):32
Bri2 BRICHOS client specificity and chaperone activity are governed by assembly state.
Chen G, Abelein A, Nilsson HE, Leppert A, Andrade-Talavera Y, Tambaro S, Hemmingsson L, Roshan F, Landreh M, Biverstål H, Koeck PJB, Presto J, Hebert H, Fisahn A, Johansson J
Nat Commun 2017 12;8(1):2081
Non-pathological amyloid in silks and the NT* solubility tag
Non-pathological amyloid with well-defined physiological roles has been identified. Spider silk shares amyloid-like properties and is one of the toughest biomaterials, making it a fascinating target for biomimicry. Spider silks are spun from large proteins (spidroins) that undergo a rapid transformation from extremely concentrated soluble dope with helical and disordered conformations to solid β-strand-rich fibers, and this process is regulated by globular N- and C-terminal (NT and CT) domains. The CT domain and also the repetitive regions have been shown to form amyloid structure; however, the importance and the underlying mechanisms of the amyloid-like formation are still poorly understood. The NT domain (NT) plays a central role to maintain the solubility, and has successfully been used to produce highly hydrophobic and aggregation-prone peptides/proteins, like Aβ42, Aβ40, SP-C, β17 and chaperones. This project aims to understand non-disease associated amyloid-like formation, and further extend the solubility tag NT* applications to tau protein and bioactive enzymes.
High-yield Production of Amyloid-β Peptide Enabled by a Customized Spider Silk Domain.
Abelein A, Chen G, Kitoka K, Aleksis R, Oleskovs F, Sarr M, Landreh M, Pahnke J, Nordling K, Kronqvist N, Jaudzems K, Rising A, Johansson J, Biverstål H
Sci Rep 2020 01;10(1):235
High intracellular stability of the spidroin N-terminal domain in spite of abundant amyloidogenic segments revealed by in-cell hydrogen/deuterium exchange mass spectrometry.
Kaldmäe M, Leppert A, Chen G, Sarr M, Sahin C, Nordling K, Kronqvist N, Gonzalvo-Ulla M, Fritz N, Abelein A, Laίn S, Biverstål H, Jörnvall H, Lane DP, Rising A, Johansson J, Landreh M
FEBS J 2020 Jul;287(13):2823-2833
A spidroin-derived solubility tag enables controlled aggregation of a designed amyloid protein.
Sarr M, Kronqvist N, Chen G, Aleksis R, Purhonen P, Hebert H, Jaudzems K, Rising A, Johansson J
FEBS J 2018 05;285(10):1873-1885
Diversified Structural Basis of a Conserved Molecular Mechanism for pH-Dependent Dimerization in Spider Silk N-Terminal Domains.
Otikovs M, Chen G, Nordling K, Landreh M, Meng Q, Jörnvall H, Kronqvist N, Rising A, Johansson J, Jaudzems K
Chembiochem 2015 Aug;16(12):1720-4
Carbonic anhydrase generates CO2 and H+ that drive spider silk formation via opposite effects on the terminal domains.
Andersson M, Chen G, Otikovs M, Landreh M, Nordling K, Kronqvist N, Westermark P, Jörnvall H, Knight S, Ridderstråle Y, Holm L, Meng Q, Jaudzems K, Chesler M, Johansson J, Rising A
PLoS Biol 2014 Aug;12(8):e1001921
Sequential pH-driven dimerization and stabilization of the N-terminal domain enables rapid spider silk formation.
Kronqvist N, Otikovs M, Chmyrov V, Chen G, Andersson M, Nordling K, Landreh M, Sarr M, Jörnvall H, Wennmalm S, Widengren J, Meng Q, Rising A, Otzen D, Knight SD, Jaudzems K, Johansson J
Nat Commun 2014 ;5():3254
Full-length minor ampullate spidroin gene sequence.
Chen G, Liu X, Zhang Y, Lin S, Yang Z, Johansson J, Rising A, Meng Q
PLoS One 2012 ;7(12):e52293
Bachelor, Master, PhD or Postdoc Project
If you are interested in doing a Bachelor, Master, PhD or Post-doc project with us and highly motivated, please contact gefei.chen@ki.se for potential projects and vacancies.
