Our research
Project 1: Mechanism of circadian transcription in the 3D nuclear architecture
Light-sensitive organisms have evolved strategies to adjust their daily activity to a 24-hour cycle of daylight and darkness. This form of adaptation to diurnal changes in the environment is regulated by endogenous biological clocks. Circadian rhythm is “entrained” or readjusted to the geophysical time every day by external time cues, such as light and food intake. Endogenous biological clocks govern our physiology, energy homeostasis and behaviour, whereas the deregulation of circadian homeostasis increases the risk of developing a wide range of multifactorial diseases, such as psychiatric, endocrine, metabolic diseases and cancer. Our research has uncovered a novel principle of circadian transcriptional regulation in the 3D nuclear architecture. We have thus shown that rhythmic mobility of circadian loci between transcriptionally permissive and repressive sub-nuclear environments facilitates the spatiotemporal coordination of circadian transcription (Fig. 1).
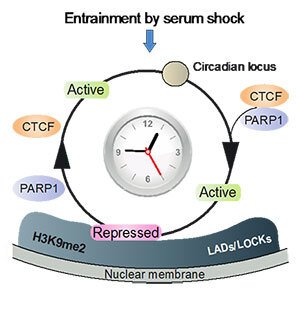
Figure 1. Circadian chromatin transitions in the 3D nuclear architecture. Entrainment by serum shock involves the coordinated and rhythmic recruitment of circadian genes from the transcriptionally permissive nuclear interior to the repressive nuclear periphery. Moreover, circadian chromatin mobility to and from the lamina is regulated by oscillating complex formation between the transcription factor CTCF and poly(ADP-ribose) polymerase 1 (Molecular Cell 2015).
Despite efforts to understand the basic principles of circadian regulation, many challenges remain. Our group is thus focusing on the following sub-projects:
- Molecular mechanisms underlying circadian chromatin transitions in the 3D architecture of the nucleus
- Reprogramming of 3D genome organization by external time cues
- Crosstalk between different regulatory layers within the cell, such as metabolism and 3D genome organization, during the circadian cycle
Project 2: Perturbed 3D genome organization in complex diseases:
Polycystic Ovary Syndrome
We are exploring the etiology of complex diseases from entirely novel perspectives. Polycystic Ovary Syndrome (PCOS) is one of the most common endocrine disorders in women of reproductive age, with severe consequences on fertility and increased risk for obesity, metabolic diseases and endometrium cancer. Our aim is to determine the cause and effect relationship between the endocrine-metabolic deregulation and the altered circadian oscillations in gene activity in PCOS, and to pinpoint novel therapeutic targets for the metabolic and malignant consequences of the disease.
Cancer
While the survival of stage 4 breast cancer patients remains particularly poor, many patients with early stage breast cancer are over-treated due to the lack of reliable prognostic markers. As chemotherapy has been implicated in the induction of cell state change towards stem-like phenotypes, paving the way for cancer evolution, there is a need to devise novel prognostic markers and to revolutionize our approach to therapy. We have formed an interdisciplinary consortium, funded by the Knut och Alice Wallenbergs Foundation, that is aiming at uncovering the role of perturbed 3D genome organisation in the emergence of immature and plastic tumor stem-like cell states that drive tumor evolution (Fig. 2), and at identifying novel therapeutic strategies that reduce the phenotypic plasticity of tumor cells. Moreover, we are examining the cause-and-effect relationships between the frequent deregulation of 3D nuclear architecture and circadian perturbations in tumors. We are thus exploring the novel avenue of thinking that cancer cells might harness the circadian system to increase their phenotypic plasticity under changing selective pressure.
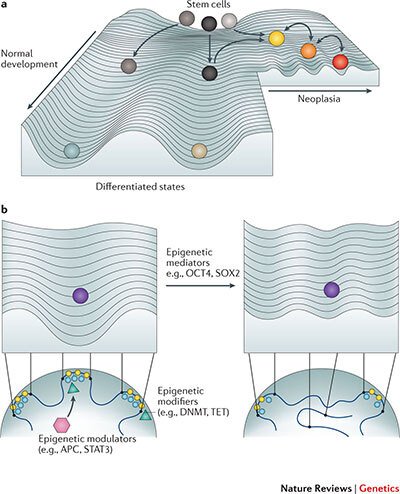
Figure 2. Perturbed 3D genome organization and tumor development. Increased phenotypic heterogeneity fuels cancer evolution under changing selective pressure on the Waddington landscape of tumor development. In a recent review we have proposed that perturbed 3D genome organization contributes to the ectopic re-activation of embryonic gene expression programs in cancer, which interferes with cellular differentiation and facilitates the emergence of stem-like immature cell states and cellular phenotypic heterogeneity (Feinberg AP, Koldobskiy M and Göndör A, Nature Reviews Genetics, 2016)
PhD courses we organise
2018 Jun: The epigenome: a platform for the integration of metabolic and signaling pathways in development and on the path to diseases
2017 Nov: The future of medicine: the role of "chance" in development, evolutionary adaptation and diseases
2017 Oct: The epigenome: a platform for the integration of metabolic and signaling pathways in development and on the path to diseases
2017 Jan: The epigenome: a platform for the integration of metabolic and signaling pathways in development and on the path to diseases
2016 May: Epigenome: the interface between nutrition and health
2015 Oct: The third dimension of nuclear functions
Vacancies
Enquiries about experimental and computational postdoctoral positions are welcome.
