Research focus
Our lab has two main lines of research: 1) mechanisms of cell fate decisions in humoral immune responses and 2) biology of innate-like lymphocytes. To learn more about some of our projects, watch our recent Global ImmunoTalks and ImmuneZoom seminars:
Mechanisms of cell fate decisions in humoral immune responses underlying the formation of immunological memory.
Antigen-dependent B cell differentiation is an unusual developmental process, in which the two main outputs of the humoral immune response – antibody-secreting plasma cells and quiescent memory B cells – can be generated by two distinct pathways: either directly from early activated B cells, or after transition through a transcriptionally distinct germinal center (GC) B cell state.
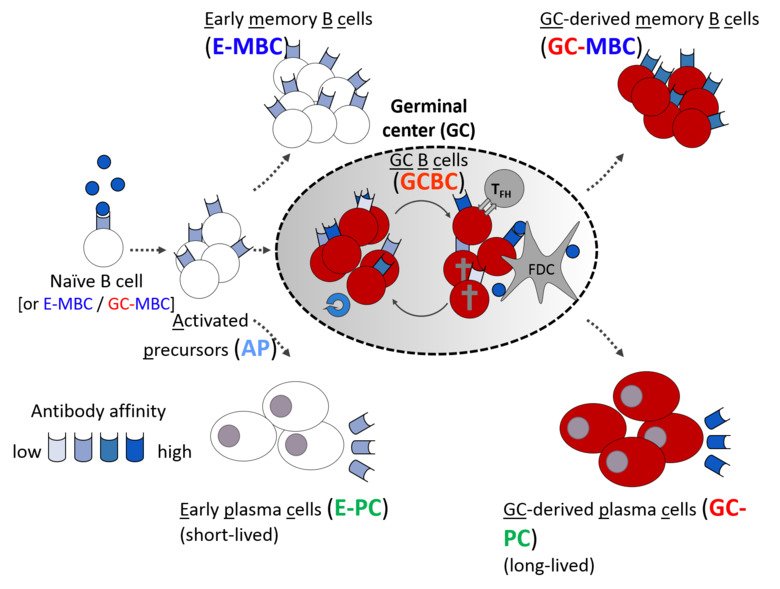
The first pathway can generate antibody-secreting cells very rapidly, providing the first line of defense against pathogens. Participation in the GC reaction allows to increase the affinity of antibodies but takes longer time. Understanding these developmental paths is of crucial importance, as the quality of immunological memory can vary greatly after infection and vaccination, resulting in a different degree of protection during the subsequent microbial challenge. We are investigating cellular and molecular mechanisms that govern formation of these distinct waves of memory B cells and plasma cells as well as their functional properties. Pursuing this goal, we have recently discovered cell intrinsic functions of the transcription factor Bhlhe40 in both B and T lymphocyte subsets involved in the GC reaction (Rauschmeier et al, J Exp Med, 2022). Moreover, our recent work also revealed that a large fraction of activated B lymphocytes in the first days of the response undergoes ‘differentiation by default’ to germinal center-independent memory B cells, a population that was often viewed as a minor, insignificant subset of B cell memory (Glaros et al, Immunity, 2021). These unexpected observations change our views on the composition of the long-term B cell memory pool and open many new questions about the formation of immunological memory to infections and vaccines that our lab is currently pursuing.
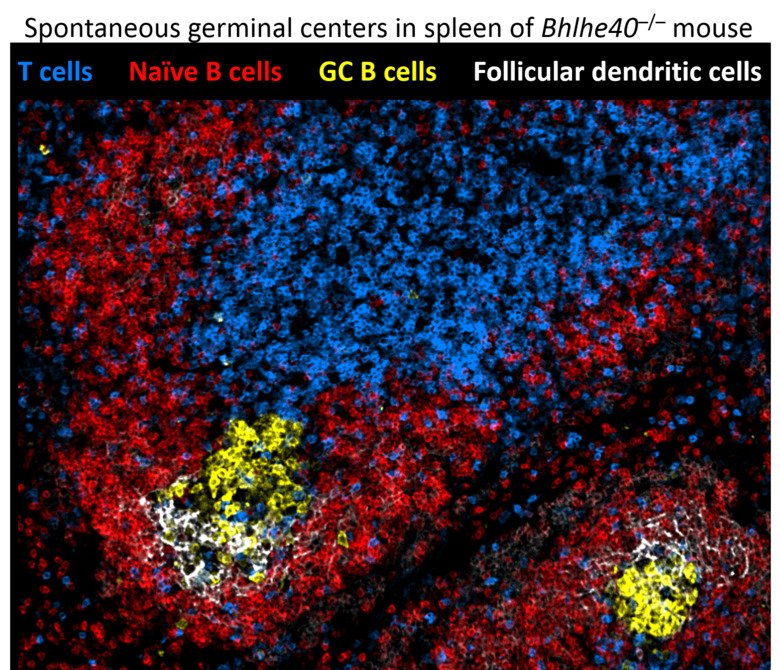
Biology of innate-like lymphocytes.
Lymphocytes of the adaptive immune system can be functionally split into two major groups. Conventional T and B cells are characterized by extremely broad antigen receptor repertoires and therefore can respond to a diverse variety of antigens, but their activation, expansion and differentiation into effector cells requires several days. In contrast, innate-like B and T cells (e.g. B1 cells, iNKT cells and many γδT cell subsets), exhibit more focused antigen receptor repertoires and are maintained in a constantly pre-activated state. These cells are thought to provide the first line of defense against pathogens and/or play a role in tissue homeostasis and repair. While we have a fairly good understanding of the biological functions, antigen specificities, selection rules and transcriptional programs for most conventional subsets, this is not the case for the majority of innate-like populations. Many basic questions that for conventional lymphocytes were solved years or even decades ago remain unanswered for these non-conventional subsets.
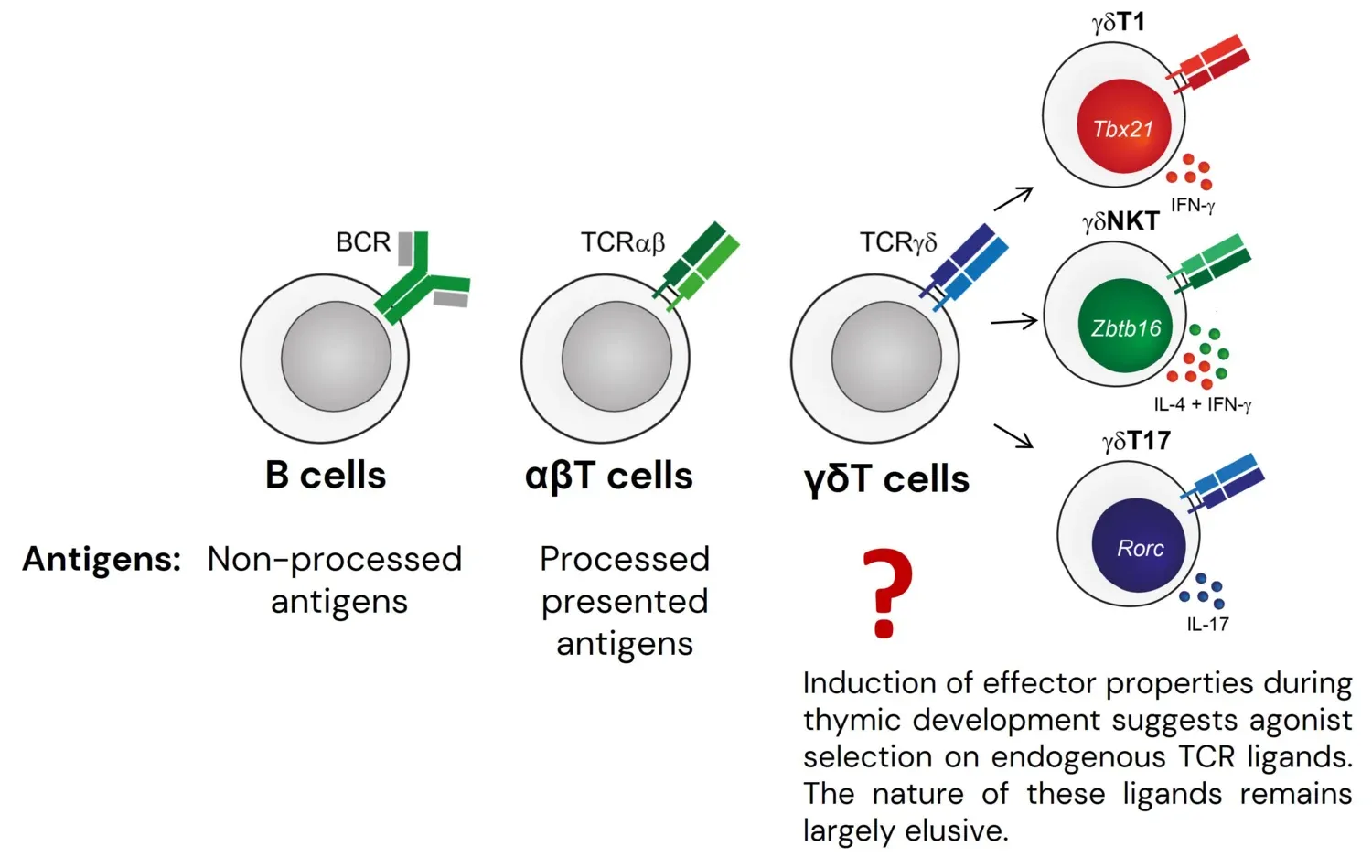
Out of the three types of lymphocytes that constitute the adaptive immune system of jawed vertebrates, γδT cells – a population particularly enriched for cells with innate-like properties – are the only cell type for which we do not understand the principles of antigen recognition and know very little about the nature of these antigens. It is believed that many γδT cells undergo agonist selection on endogenous ligands, which results in the acquisition of innate-like effector properties. In line with agonist selection of γδT cells, our results demonstrate that a large fraction of γδT cells recognizes a diverse set of unknown self antigens. Strikingly, we showed that the majority of these antigens do not belong to known families of γδTCR ligands, indicating that the known endogenous γδTCR ligands represent just the very tip of the iceberg, and that many innate-like γδT cells are likely to undergo agonist selection on a diverse set of unidentified molecules that do not belong to known families of γδTCR ligands. This result highlights how little we understand about the biology of these cells. To start filling this gap in understanding we established a screening pipeline that already allowed us to identify several novel γδTCR ligand candidates. We are now exploring the functional consequences of this recognition in in vivo models. The pipeline for ligand identification that we had established should allow us to find γδTCR ligands across a variety of physiological and pathological contexts and we are currently applying our approach to identification of antigens for γδTILs in human cancers.
In addition to our interest in the antigen receptor specificities of innate-like T cells, we are also interested in their functions. It was recently shown that many innate-like T cells in the thymus represent tissue-resident cells rather than newly developed cells about to egress to the periphery. The reasons for the accumulation of effector T cells at the site of T cell development, where pathogen encounter is unlikely, remained unclear. We have demonstrated that thymic innate-like T cells, a population that express molecules characteristic for all major activated T cell subsets, can induce highly efficient CD8 T cell tolerance to inflammation-associated self-antigens, a class of self-antigens that at the periphery largely mirror the spatial and temporal distribution of pathogen-derived molecules (You et al, Nat Immunol, 2024). As the thymus harbors a complex constitutively active inflammatory network, with thymus-resident innate-like T cells representing one of its central nodes, these results suggest that this constitutive expression of inflammation-associated molecules in the thymus may have evolved to ensure the induction of tolerance to this unique class of self-antigens.

