Our Research
Until recently, only snapshots of molecules could be observed, hiding their mode of operation. We employ Nuclear Magnetic Resonance (NMR), and other biophysical techniques, to investigate the molecular mechanism of RNA function. When function of these molecular machines becomes apparent, it also provides a variety of unique new drug targets. The lab develops methods in NMR and RNA biochemistry to address these questions. Current projects include viral, bacterial and eukaryotic regulatory RNAs, e.g. microRNAs, ribosomal RNAs or RNA from HBV.
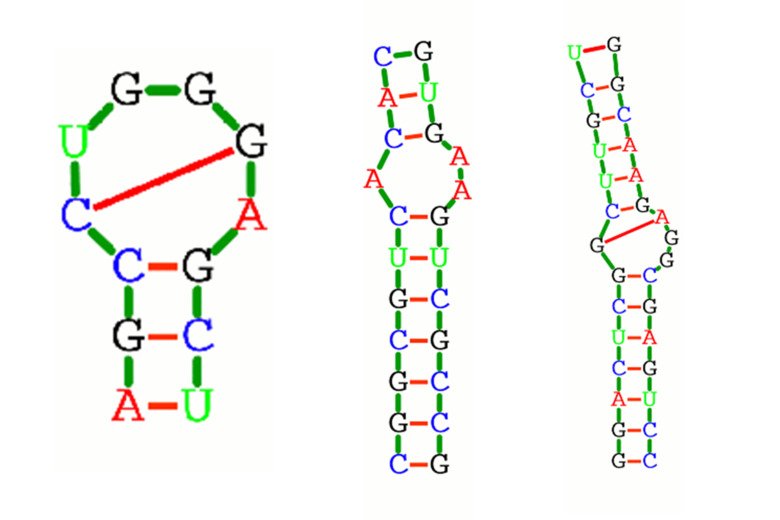
How RNA move – excited states of RNA (Nature 2012):
News
2018
- The Petzold Group moves into KI's new building Biomedicum.
- Katja receives the AMPERE Prize for Young Investigators and will give a prize lecture at this years Euromar in Nantes.
- Our method development paper “Efficient detection of structure and dynamics in unlabeled RNAs: the SELOPE approach” is accepted to Chemistry. Congratulations to Judith, Emilie and Hampus for a great work!
- Analytical and Bioanalytical Chemistry. Congratulations to Lorenzo, Hampus and Maja for a great publication!
- Our review article “A guide to RNA large scale sample preparation” is accepted into Analytical and Bioanalytical Chemistry. Congratulations to Lorenzo, Hampus and Maja for a great publication!
2017
- Hannes Feyrer's work was promoted to talk at the meeting of the RNA Society of Sweden, Lund.
- Lorenzo Baronti has received the Suraj Manroa Student Poster Price for presenting his work at the Euromar 2017.
- Lorenzo Baronti has been selected to attend the EMBO practical NMW course in Basel this summer in a strong competition.
- Katja has been invited to become a Faculty 1000 member.
- Congratulations to Judith, who received the highly competitive Marie Curie Fellowship.
2016
- Katja received funding from the Foundation for Strategic Research (SSF) for five years.
- Katja received the Sven och Ebba-Christina Hagbergs Pris for her work on RNA structure and function using NMR. This price is given to young group leaders who have shown potential.
- Katja and Emma receive the Knut och Alice Wallenberg Project grant. Our project on miRNA targeting got funded by the prestigious Wallenberg foundation – Thank you!!
- Congratulations to Emilie (first author), Judith and our collaborator Patrik Lundström for their great work to extend R1ρ timescales and probes with 1H R1ρ.
- Our paper “Visualizing Transient Watson-Crick Like Mispairs in DNA and RNA Duplexes” Nature, 2015, is as of March/April 2016 a highly cited paper (received enough citations to place it in the top 1% of its academic field) based on a Web of Knowledge/ISI.
- Master student Karen Schriever finishes her project in the Petzold lab.
- Master student Luís Silva will join the Petzold lab for a one year project.
- Congratulations Luís Silva on receiving an Erasmus scholarship for his Master Thesis!
- Lorenzo Baronti received the travel grant from the Biophysical Society for the “CHIANTI WORKSHOP ON MAGNETIC RESONANCE FOR CELLULAR STRUCTURAL BIOLOGY” for his work on miRNA targeting.
- During 2016, Bachelor student Noah Hopkins joins the Petzold lab for a project.
- Postdoc Judith Schlagnitweit and Master student Karen Schriever joins the group.
- The Petzold Lab co-organizes the workshop in Stockholm June 12-15, where experts in RNA biology and RNA structure meet
Available positions
Applications for Master Students are welcome in the field of NMR or RNA biochemistry. Students have preferably a background in biochemistry, chemistry, molecular biology or physics with an interest in inter-disciplinary research.
We are always looking for talented Master, PhD students and Postdocs. Please send a 1 page coverletter, 2 page CV, publication record and list of references to Katja Petzold.

