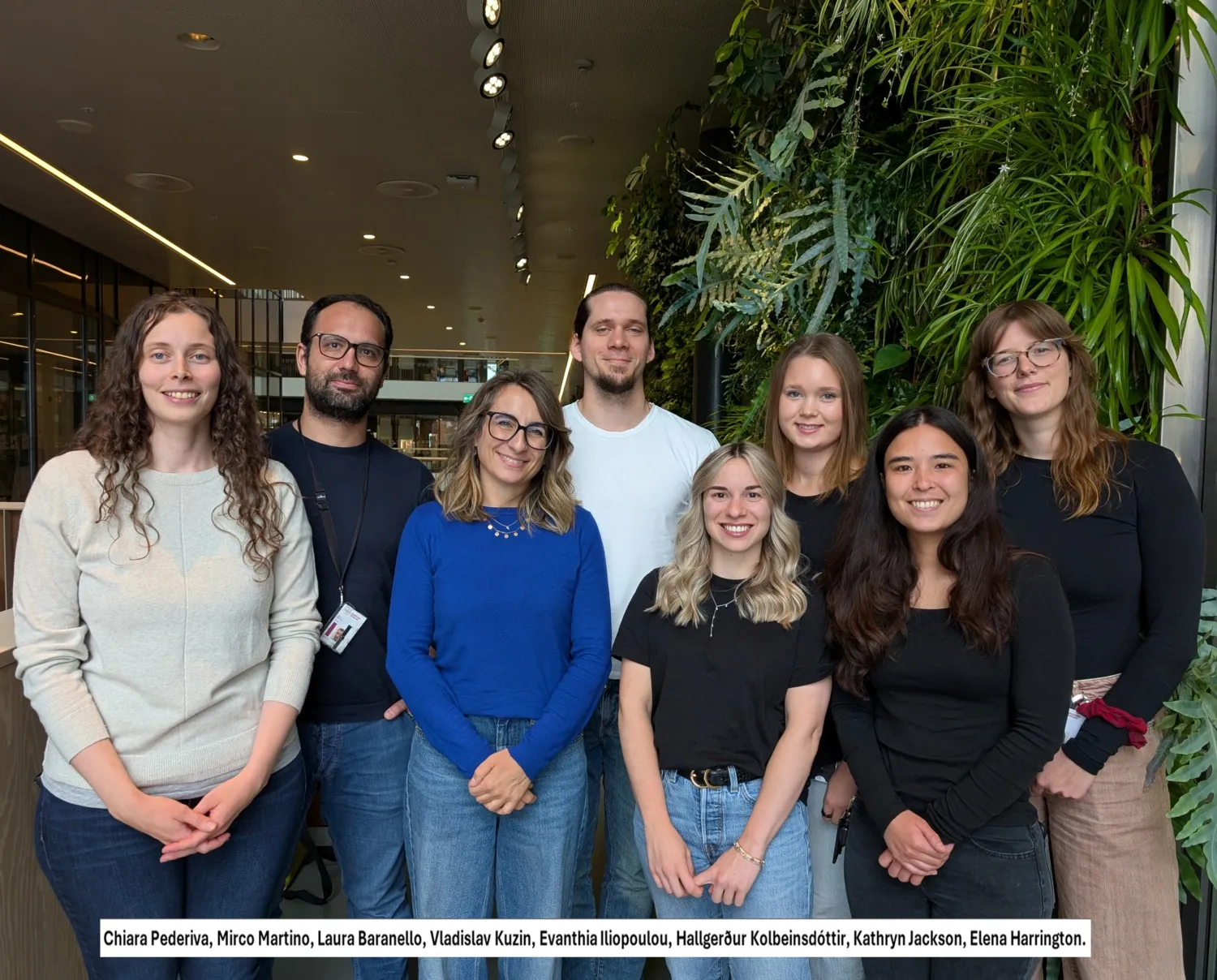
Our research
Cancer is a biologically complex disease that causes significant deaths in the human population. Pharmaceuticals that inhibit enzymes called topoisomerases are effective at killing many types of cancer cells. Unfortunately, the body’s healthy cells are also damaged by this treatment. Development of tumor-specific topoisomerase inhibitor-based therapies will require better knowledge of the mechanism of topoisomerase activity.
Topoisomerases are important cellular enzymes; they are involved in processes in which genes are copied, or when DNA is replicated prior to cell division. They unwind the DNA double helix, so that the enzymes that are going to transcribe genes or replicate DNA strands are able to do so.
Although conventionally considered to be constitutively active enzymes, recent evidence show that topoisomerases execute their function through regulatory interactions with partner proteins that modulate their activity to affect the transcriptional outcome. Understanding the mechanism of this regulation might provide a new strategy to affect topoisomerase activity in cancer cells.
Recently, the Baranello group has identified a unique complex comprised of MYC, TOP1 and TOP2A named the “topoisome”, in which MYC stimulates the topoisomerase’s activity to enhance transcription and replication in cancer cells (Molecular Cell, 2022). Targeting cancer-specific properties, such as the topoisome, could result in highly selective cancer therapeutics with minimal off target/side effects.
Projects
Our ongoing and future investigations are based on these findings. We specialise in a variety of approaches including biochemical assays, next-generation sequencing techniques and bioinformatics, genome editing and drug screens to:
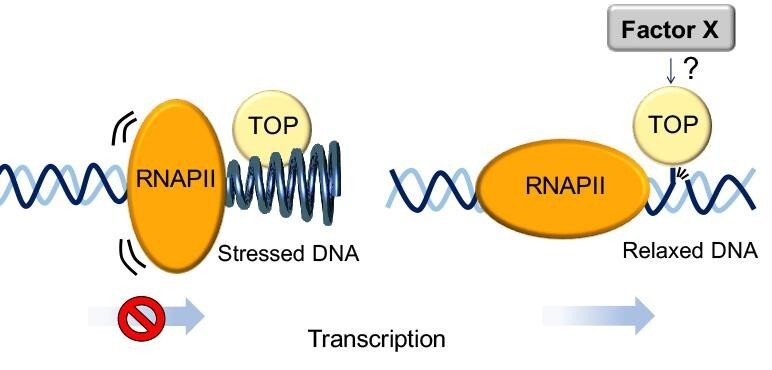
- Determine how the topoisome coordinates with RNA polymerase II to amplify transcription and enhance MYC-driven lymphomas
- Identify new proteins regulating topoisomerase 1 and topoisomerase 2 activity. Among the potential partners we focus on transcription and chromatin factors.
- Understand the molecular details of how topoisomerases are regulated by their protein partners during transcription.
- Identify drugs targeting the stimulation of topoisomerase activity in cancer cells.
If you are interested in our research
Collaborations
Our research is performed with a number of national and international collaborators:
- Prof. Björkegren, Karolinska Institutet, Sweden
- Dr. Levens, National Institutes of Health, USA
- Associate Prof. Mazza, San Raffaele Hospital, Italy
- Prof. Dr. Nishiyama, Nagoya University, Japan
- Associate Prof. Oberdoerffer, John Hopkins, USA
- Prof. Dr. Shirahige, Tokyo University, Japan
- Prof. Dr. Westerberg, Karolinska Institutet, Sweden
Conference organiser
EMBO workshop “DNA Topology in genomic transactions
Cell and Molecular Biology seminar series
Co-Organizer of Cell and Molecular Biology Departmental Retreat 2023