Our research
We have uncovered highly relevant connections between (i) cilia, ciliary genes and cilia-based signalling and (ii) (candidate genes for) different human brain conditions or disorders, like dyslexia or schizophrenia. The biological pathways behind these brain phenotypes are largely unknown. And our understanding of neuronal cilia is still rudimentary.
Fundamental questions about ciliary involvement in brain development, function and behavioural output in normal and disease states remain. What are the mechanisms by which cilia orchestrate cellular signalling in neuronal development? When and how does ciliary signalling control cell fate during expansion of the neural progenitor cell pool and their differentiation and organisation into neurons during brain development? Addressing these questions is crucial for understanding disease aetiology and for eventually developing treatment regimens for brain disorders.
Our work lets us hypothesise that proper cilia function impacts differentiation of neural progenitor cells and early-stage neurons, including polarisation and neurite outgrowth, and thereby neuronal migration and circuit formation later on. Different non-lethal cilia malfunctions may thus cause different brain phenotypes (or disease states) depending on the affected neuron type and its location in the brain.
We address these hypotheses by:
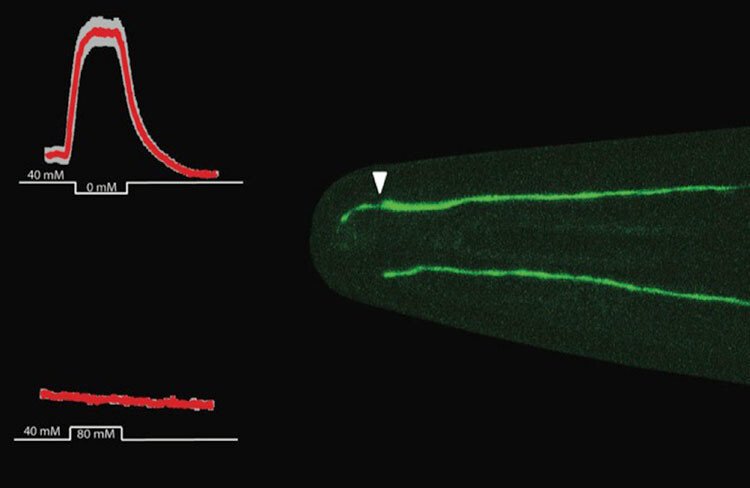
- Determining in cultured human neurons and a whole-animal model, the worm C. elegans, causative connections between cilia formation, structure and function and aspects of neuronal development.
- Analysing transcriptomes of differentiating (human) neurons, including bioinformatics-based cross-correlations with ciliary gene lists and human candidate genes for various brain phenotypes.
- Analysing in the worm C. elegans behavioural output of disease-associated mutations in evolutionarily conserved (human and worm) ciliary genes connected to brain phenotypes.
Our goal is to create proper spatiotemporal information about ciliary localisation and function of proteins encoded by disease genes to better understand human brain development in the context of neural progenitor cells, and differentiating and mature functional neurons.
Research networks
- KI Neurosciences network (StratNeuro)
- European C. elegans researcher network (EU COST Action Network BM1408)
- Swedish-Korean research cooperation network (VR and STINT, Korean NRF)
