About our research
We are dedicated to studying the molecular mechanisms regulating normal physiology as well as disease pathogenesis. We do this by combining advance animal models with genetic and molecular approaches. Our research is multidisciplinary and comprehensive, going all way from genes and molecules to cells, genetically engineered mice and preclinical studies.
We primarily use loss-of-function mutant mice to reveal the role of genes in disease mouse models, and combine them with cellular and molecular techniques, such as flow cytometry, histology, molecular biology and cell culture. We also use ex vivo specialized primary cultures, including 3D organoids, virus-based gene editing, and omic technologies.
We focus our studies in the fields of cancer, immunology, intestinal biology as well as cardiometabolic diseases, with a particular molecular interest for exploring how the process of protein ubiquitination participate in physiology and disease pathogenesis.
Ongoing research
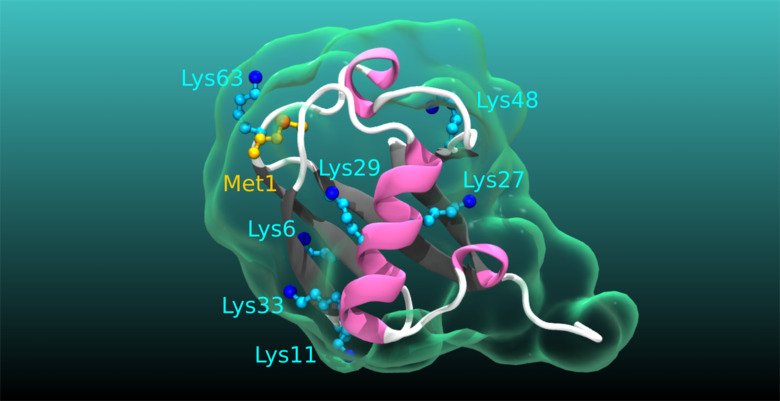
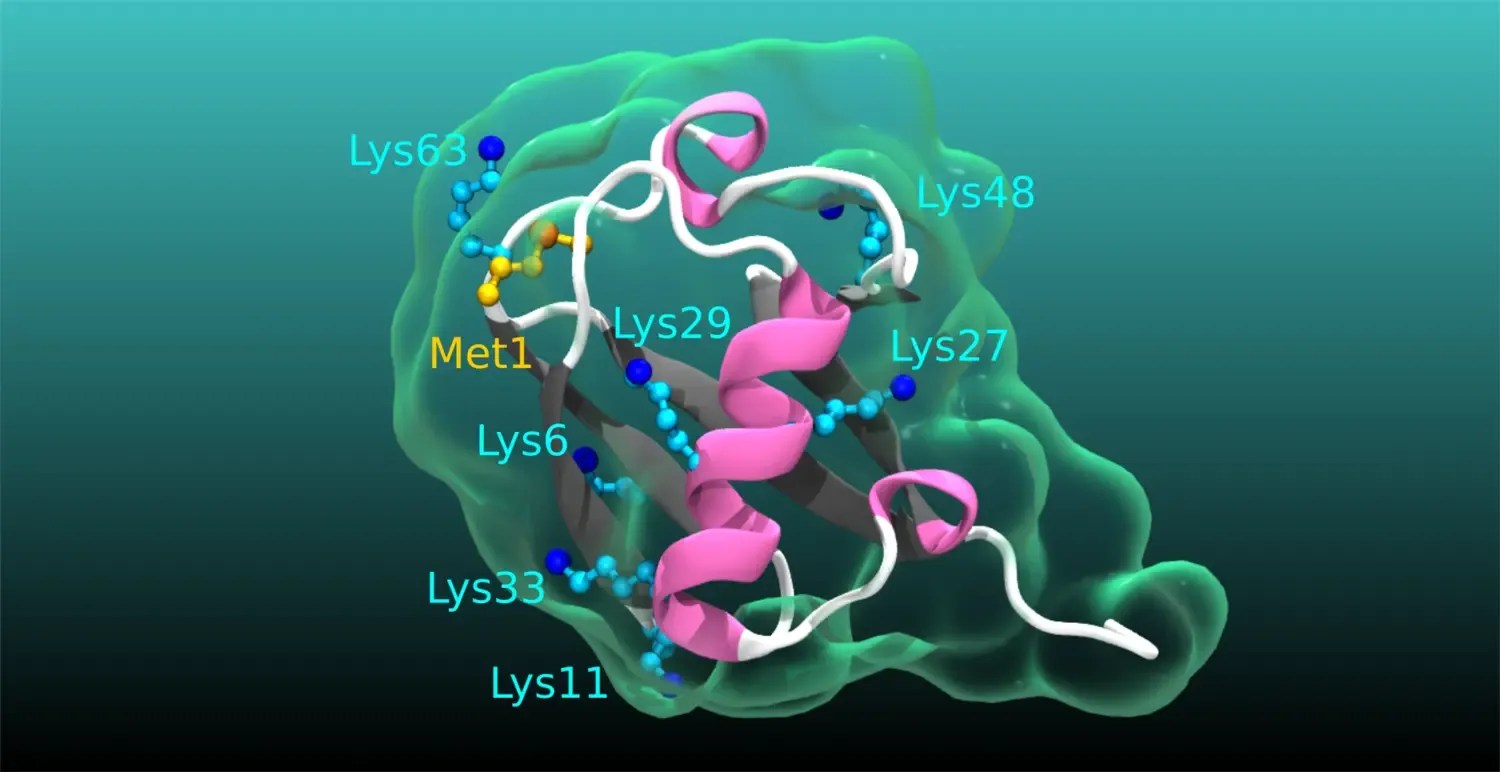
Ubiquitination in physiology and disease: Even though numerous human pathologies are linked to alterations in the process of ubiquitination, an essential posttranslational protein modification, we still do not fully understand the true complexity, specificity, and dynamics of the ubiquitin code. Specifically, the cellular and physiological processes regulated by many deubiquitinating enzymes, the enzymes that remove ubiquitination, as well as the pathophysiological roles of atypical ubiquitin chains, chains of ubiquitin linked by lysines (K) K6, K27, K29, K33, remain obscure. Our current main research aims to narrow these knowledge gaps. We are undertaking diverse projects to revealing the functions and impact of atypical ubiquitination and the related deubiquitinases in intestinal homeostasis, cancer, immune related disorders and cardiometabolic diseases.
Gender dimorphism: We recently revealed how the thymus stroma adapts to integrate the hormonal, immune and metabolic responses of pregnant females mice, safeguarding via the immune system the development of the embryos (Nature 2021). We keep interested in this line of research and have also projects aiming to better understand gender dimorphism in relation to immune and cardiometabolic diseases.
The significance of our research
Comprehensive preclinical studies using model organisms are essential to identify new disease-related genes, better understand the complexity and etiology of diseases and can also serve as platforms for drug discovery programs. Our integrative approach to use functional genetics analyses to provide thorough in vivo and mechanistic insights will increase our molecular understanding of the pathogenesis of diseases and could set the molecular basis for the design of novel strategies for therapies.