Team Immuno-metabolic reprogramming during viral infection
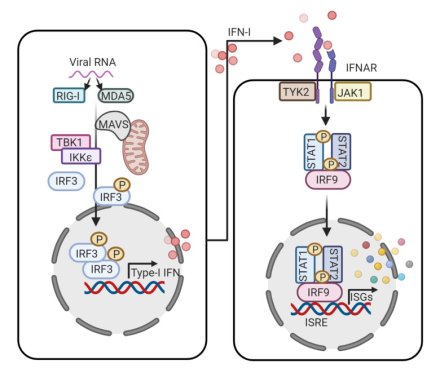
Our projects investigate immuno-metabolic reprogramming during viral infections, focusing on how viruses hijack host metabolism to facilitate replication and development of host-directed therapy. Using multi-omics systems biology, metabolic modeling, and experimental assays, we aim to identify key metabolic and signaling alterations induced by viral infections.