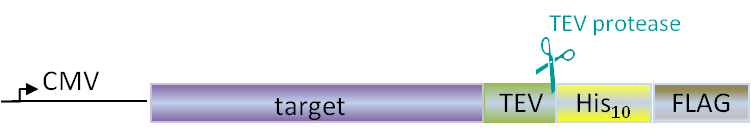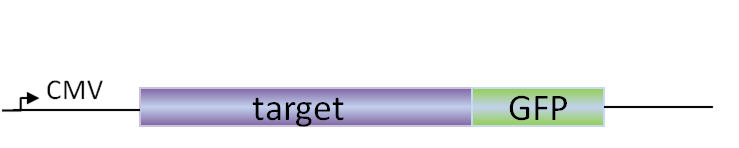Protein Production Platform
The PSF offers services in protein production, molecular biology, and small-scale expression tests. We offer protein expression services in E. coli as well as eukaryotic cells.
BACTERIAL PROTEIN EXPRESSION
E.coli expression cultures are typically 1.5-4.5 liters, depending on the expected yield. Cultures are typically grown in TB or LB at 37°C before down-tempering to 18 °C. After induction protein production is alowed to continue over night.
High throughput protein purification
Harvested cells are usually stored at -80°C before lysis by sonication, clarification by centrifugation and filtering prior to applying the sample to an ÄKTAxpress system for a two-step purification. Our standard purification scheme consists of an IMAC affinity step utilizing the His-tag followed by a size exclusion chromatography step (SEC, gel filtration). Other purification protocols and polishing steps are available and adjusted to the project needs. For expression we mostly use BL21(DE3) T1R pRARE2, but several other strains are used when necessary. The purity and quality of the purified protein is assessed by SDS-PAGE, gel filtration elution profile and in cases MS data.
Small scale expression and solubility test
Small scale expression test where the levels of total protein as well as the yield after affinity purification are assessed by SDS-PAGE analysis. Typically, 1 ml cultures are grown in 96 well blocks in Terrific Broth with expression at 18°C and over night. Harvested cells are lysed and clarified supernatants are subject to an IMAC affinity step. For expression we mostly use BL21(DE3) T1R pRARE2, but several other strains are used when necessary.
EUKARYOTIC PROTEIN EXPRESSION
PSF offer protein expression services in eukaryotic systems, predominantly HEK293 cells.
For cytoplasmic expression, we prefer to work with GFP-fusion proteins, which helps monitoring the transfection and expression efficiency. Streptags are the preferred affinity tag for purification, as Ni-resins bind many contamiting cytoplasmic proteins.
Secreted proteins are purified from the clarified media. His-tags and IMAC purification works well, but Strep-tag is also an option.
For new projects we normally run 200 ml test expression cultures. Expression and solubility are evalueted before decission is made on the continuation of a project.
Production cultures are typically 1-5 liters, with capacity to run up to 20 liters.
As a core facility with limited staff and many clients, we regretfully cannot take on projects that demands extensive development work and trouble shooting. Such projects are better suited for a doctoral student or post doc.
Full-service in eukaryotic protein expression is available for KI groups. Requests from other universities are encouraged to contact the Protein Production Sweden national infrastructure.
Cloning and high throughput cloning
As the chemical synthesis of DNA is increasingly cheap to order today, we often order the expression constructs we need. We are happy to help your clients with both design and ordering of plasmids.
PSF have long experience in efficient HTP cloning work in 96-well format. For this we have a long list of LIC (ligation independent cloning) prepared vectors, but also work with Gibson assembly protocols. The methods are robust and enables parallel cloning of many constructs in a 96 well format. Our most frequently used cloning vector is pNIC28-Bsa4 which adds an N-terminal His-tag followed by a TEV protease cleavage site to the target protein. See list of selected available vectors below.
TERMS OF SERVICE
PSF’s protein production operates on a fee-for-service basis where the academic user fee is comprised of consumables, instrument costs, university OH and part of staff and rent. The rest is covered by Karolinska Institutet and other funders such as VR. Non-academic users fees cover the full cost.
Note that the fees are charged for agreed work and not for the resulting material. The standard protocols used by PSF are developed to be the best for soluble proteins in general, but might not be suitable for any specific protein. Also, yields vary between purifications, even for proteins that are documented to behave well. Therefore, PSF leaves no guarantee that the above-described work will result in any specific amount of deliverable material. However, if an experiment fails due to technical or human error at PSF it will be repeated with no extra charge.
Projects/samples that fail during technically successful experiments are transferred back to the user together with generated data. Additional investigation or problem solving can be performed after discussion and agreement with the user.
Extensive work requests, filling up all available slots, might be divided into smaller parts in order not to block the project flow and delay delivery times for other users.

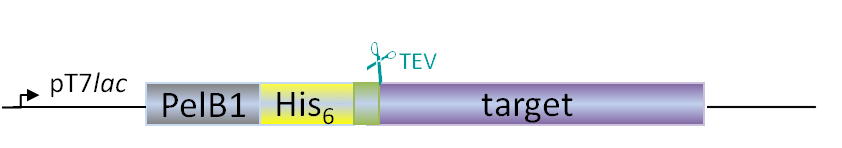
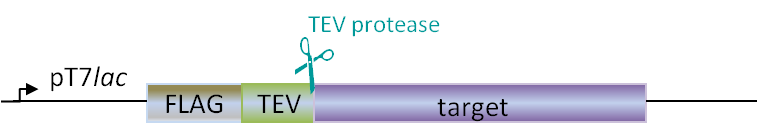
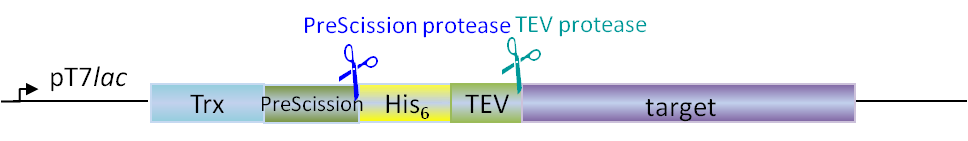
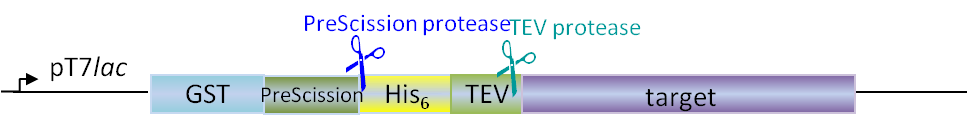
Plasmids for E. coli
Our in-house plasmid collection contains vectors for protein production with different fusion proteins in different hosts. See a list of example plasmids below. Any other plasmids needs can be solved. Feel free to contact us for questions!
-


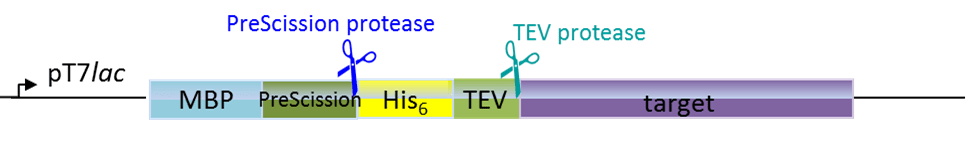
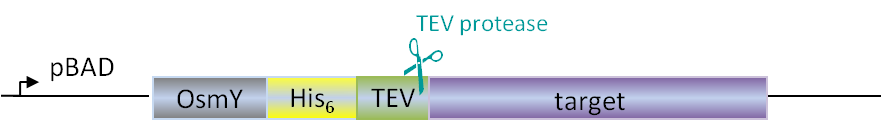












-
Plasmids for other hosts