Project: Computational and toxicological modeling of cellular metabolism
A general description of effects of toxic compounds in mammalian cells is facing several problems. Firstly, most toxic compounds are hydrophobic and partition phenomena strongly influence their behaviour. Secondly, cells display considerable heterogeneity regarding the presence, activity and distribution of enzymes participating in the metabolism of foreign compounds i.e. bioactivation/biotransformation. Thirdly, cellular architecture varies greatly.
Taken together, complexity at several levels has to be addressed to arrive at efficient in silico modelling based on physicochemical properties, metabolic preferences and cell characteristics. In order to understand the cellular behaviour of toxic foreign compounds we have developed a mathematical model that addresses these issues. In order to make the system numerically treatable, methods motivated by homogenization techniques have been applied. These tools reduce the complexity of mathematical models of cell dynamics considerably thus allowing to solve efficiently the partial differential equations in the model numerically on a personal computer.
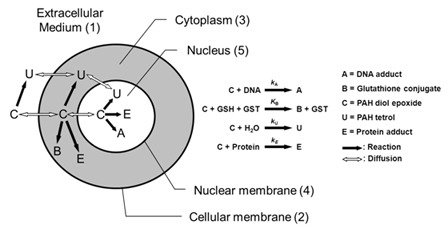
Our published results concerning metabolism and chemical solvolysis of a polycyclic aromatic hydrocarbon carcinogen show good agreement with results from measurements in V79 cell culture. Furthermore, lipophilicity was identified as an important parameter in both metabolism and formation of DNA adducts.
This project is in collaboration with Michael Hanke and his group at NADA, KTH.
Contacts
Kristian Dreij
Senior LecturerRalf Morgenstern
Professor EmeritusFinancing
- Vetenskapsrådet (NT)
Selected publications
Surface reactions on the cytoplasmatic membranes – Mathematical modeling of reaction and diffusion systems in a cell.
Chaudhry QA, Hanke M, Morgenstern R and Dreij K.
J Comput Appl Math 262, 244-260, 2014.
In silico modeling of the intracellular dynamics of polycyclic aromatic hydrocarbons.
Dreij K, Chaudhry QA, Zhang J, Sundberg K, Jernström B, Hanke R and Morgenstern R.
Tox Letters 211, suppl., S60-61, 2012.
A method for efficient calculation of diffusion and reactions of lipophilic compounds in complex cell geometry.
Dreij K, Chaudhry Q, Jernström B, Morgenstern R, Hanke M
PLoS ONE 2011 ;6(8):e23128
