Chemical Proteomics Core Facility
The Chemical Proteomics facility (ChemProt) is the national facility expert on supporting drug discovery and development by proteome-wide deconvolution of targets and mechanism of action (MoA) of small molecules or any active compounds.
ChemProt is the reference facility for chemical proteomics within SciLifeLab and BioMS national infrastructures.
The main goal of ChemProt is to decipher proteome signatures induced by an active compound for MoA elucidation and target identification. For these aims, ChemProt has adopted the latest mass spectrometry(MS)-based proteomic technologies and developed new high-throughput (HT) methods.
ChemProt also provides the HDX-MS (hydrogen-deuterium exchange mass spectrometry) technology for protein structure analysis on purified proteins with the goal of mapping the site of interaction with active compound of conformational changes in response to the applied system perturbation.
Using multidisciplinary competences and resources, ChemProt represents an innovative facility model offering complete pipelines from perturbations in cell cultures, to MS-based proteomics and data analysis, including data bioinformatics processing, publication figures and data representation.
ChemProt has recently developed high throughput (HT) chemical proteomics methods which outperforms the previous methods for target deconvolution of at least tenfold and decreases by the same factor the time of analysis of the sample amount needed for each biological replicate. The novel methods can use a high level on multiplexing for several biological replicates, up to 16 per experiment according to the latest state of the art for quantitative proteomics. Different compounds or more than one biological system can be tested simultaneously.
The LC-MS instrument park has been renewed for better supporting the new high throughput chemical proteomics.
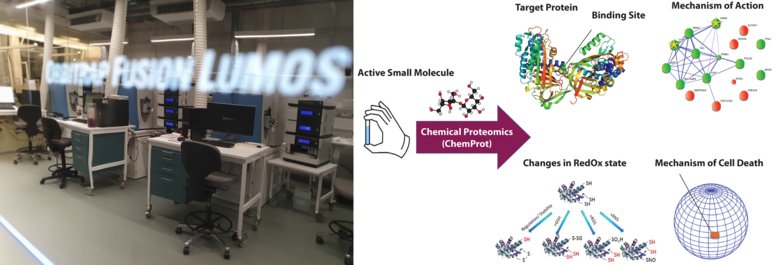
Services
ChemProt makes use of proteome-wide orthogonal approaches to find out how small molecules work when incubated in cell cultures and in lysates. Most approaches can be provided with no molecular modification for chemical probe engineering. ChemProt can provide full pipelines reproducing the biological treatments of the phenotype of interest or operating the treatment planned with the users. ChemProt also offers hydrogen-deuterium exchange mass spectrometry (HDX-MS) for mapping and characterizing the target binding site, and monitoring other structural changes. Our outcomes include full data analysis, high confidence identification of most reproducible and treatment-specific MoA signatures and target candidates. Follow up support for consequent experimental and publication purposes are provided; consultation is always provided on request and free of charge, also to guide further planning.
Summary of Current Technologies and Services
A) Chemical Proteomics for Target Discovery and Determination of Mechanism of Action (MoA) and Off-Target landscape
A1) Orthogonal methods of deep mass spectrometry-based proteomics based on
- protein solubility/stability alteration signatures in treated cell cultures and lysates;
- treatment-specific expression/degradation proteome signatures;
- changes in proteome redox balance in treated cell cultures.
Matches over two orthogonal approaches dramatically increase the chances to find the correct target. These methods are here listed (the first three are developed in-house):
- Proteome Integral Solubility Alteration (PISA) Assay, high throughput method based on protein solubility/stability upon thermal treatment (J Proteome Res 2019, DOI: 1021/acs.jproteome.9b00500) with no need of chemical engineering of compounds (now up to 16-plex with TMTpro 16-plex, i.e. 16 biological samples in one multiplex analysis);
- FITExP, based on compound-specific proteome responses (Sci Rep 2015, DOI: 1038/srep11176) and with no need of chemical engineering of compounds (now up to 16-plex with TMTpro 16-plex)
- ProTargetMiner, high throughput method to study anticancer compounds based on FITExP-derived database of proteome signatures using a library of anticancer molecules (DOI: 1038/s41467-019-13582-8);
- Identification of interactions and protein complexes after affinity-based approaches using chemical engineered probes (now up to 16-plex with TMTpro 16-plex)
- Thermal Proteome Profiling (TPP), based on protein melting curve fitting and Tm extrapolation (Nat Commun 2019, DOI: 1126/science.1255784)
- RedOx Proteomics, for specific proteome changes in the reduction-oxidation balance
A2) “System-wide Identification and prioritization of Enzyme Substrates by Thermal Analysis” (SIESTA), a new method developed in-house (Nat Commun 2021, DOI: 10.1038/s41467-021-21540-6) for identification of enzyme protein substrates. It relyes on the enzymatic post-translational modifications of substrate proteins changing their solubility and it assesses solubility changes of proteins in cell lysate when a recombinant protein and a co-factor are simultaneously added to the lysate in excess.
A3) Additional proteomics services related to / coordinated with the above (e.g. PTMs, deep quantitative proteomics, etc.)
B) Protein Structural Analysis using Hydrogen-deuterium exchange mass spectrometry (HDX-MS)
HDX-MS workflows are optimized at ChemProt for:
B1) Binding site mapping and characterization for: protein-small molecule interactions; protein-peptide interactions; protein-protein interactions, including epitope mapping;
B2) Conformational changes monitoring to study effects of mutations, to analyze misfolding and for biosimilar characterization
Equipment
ChemProt laboratories are at located at Biomedicum, Karolinska Institutet. ChemProt represents an innovative facility model offering complete pipelines and including different laboratories for cell culture, sample preparation, LC-MS and bioinformatic analysis.
Cell culture lab Instruments: Two laminar flow cabinets, Two CO2 incubators, Two Microscopes, Eppendorf centrifuge, BioRad cell counter, etc.
Centrifuges: Beckmann XPN-80 ultracentrifuge, Speedvac centrifuges , Eppendorf centrifuges for large and small tubes
Mass Spectrometers:
- Two Orbitrap Fusion Lumos, Thermo Scientific
- Two Orbitrap Q Exactives HF-X, Thermo Scientific
- One Orbitrap Fusion, Thermo Scientific
- Two Orbitrap Q Exactives HF, Thermo Scientific
- One Orbitrap Q Exactive, Thermo Scientific
UPLC:
- Two Ultimate 3000 RSLCnano for double column operation
- Five UltiMate 3000 RSLCnano / nEasy-LC 1000 for nLC-MS
- Ultimate 3000 for off-line high pH fractionation;
- Three UltiMate 3000 Nano LC or nEasy-LC 1000 for nLC-MS
Data Analysis Harware and Software:
Four PC stations, Proteome Discoverer 2.4, BioPharma Finder, Peaks, MaxQuant, Mascot, SIMCA, GraphPad PRISM, Excel, R, etc.
Recent projects
System-wide identification and prioritization of enzyme substrates by thermal analysis.
Saei AA, Beusch CM, Sabatier P, Wells JA, Gharibi H, Meng Z, Chernobrovkin A, Rodin S, Näreoja K, Thorsell AG, Karlberg T, Cheng Q, Lundström SL, Gaetani M, Végvári Á, Arnér ESJ, Schüler H, Zubarev RA
Nat Commun 2021 02;12(1):1296
Capillary leakage provides nutrients and antioxidants for rapid pneumococcal proliferation in influenza-infected lower airways.
Sender V, Hentrich K, Pathak A, Tan Qian Ler A, Embaie BT, Lundström SL, Gaetani M, Bergstrand J, Nakamoto R, Sham LT, Widengren J, Normark S, Henriques-Normark B
Proc Natl Acad Sci U S A 2020 12;117(49):31386-31397
Lysine-specific demethylase 1A restricts ex vivo propagation of human HSCs and is a target of UM171.
Subramaniam A, Žemaitis K, Talkhoncheh MS, Yudovich D, Bäckström A, Debnath S, Chen J, Jain MV, Galeev R, Gaetani M, Zubarev RA, Larsson J
Blood 2020 Nov;136(19):2151-2161
Thermal Proteome Profiling Identifies Oxidative-Dependent Inhibition of the Transcription of Major Oncogenes as a New Therapeutic Mechanism for Select Anticancer Compounds.
Peuget S, Zhu J, Sanz G, Singh M, Gaetani M, Chen X, Shi Y, Saei AA, Visnes T, Lindström MS, Rihani A, Moyano-Galceran L, Carlson JW, Hjerpe E, Joneborg U, Lehti K, Hartman J, Helleday T, Zubarev R, Selivanova G
Cancer Res 2020 04;80(7):1538-1550
ProTargetMiner as a proteome signature library of anticancer molecules for functional discovery.
Saei AA, Beusch CM, Chernobrovkin A, Sabatier P, Zhang B, Tokat ÜG, Stergiou E, Gaetani M, Végvári Á, Zubarev RA
Nat Commun 2019 12;10(1):5715
Proteome Integral Solubility Alteration: A High-Throughput Proteomics Assay for Target Deconvolution.
Gaetani M, Sabatier P, Saei AA, Beusch CM, Yang Z, Lundström SL, Zubarev RA
J Proteome Res 2019 11;18(11):4027-4037
Using the Facility
Please feel free to contact us (massimiliano.gaetani@ki.se) and come visit us at KI Biomedicum. You will receive consultation free of charge, finding out about feasibility, details and better solutions for your project or your potential one.
From the 8th of April 2021 Chemical Proteomics is available under the iLab system for KI core facilities. Our users are required to log in iLab , and submit a project request to Chemical Proteomics. Please contact us for any questions.
ChemProt is also the national facility expert of Chemical Proteomics within the Swedish infrastructure for biological mass spectrometry (BioMS). ChemProt is responsible for the entire Chemical Proteomics technology and also provider of protein structural analysis through HDX-MS. Through BioMS and SciLifeLab we can support your project and cover relevant costs of it by national funding. The users will be instructed on how to register services to our facility also through an application at the BioMS portal for the "Chemical Proteomics” technology.
Guidelines
From October 1, 2020 new guidelines are applied, in line with the guidelines introduced by KI on 1/1/2018
After an initial discussion of the project and before sample submission, the user is kindly requested to provide a brief description of the project and for that provided with a project description template and necessary help for its completion. The final project description will undergo internal facility approval and necessary suggestions to optimize the project design and maximize data quality and scientific impact of the facility application in the project.
The user could choose different levels of services, summarized below:
- Contract/service analysis: Research data acquisition. No data analysis or interpretation will be provided by facility. Archiving and public access is customer’s responsibility. No intellectual property will be retained by the facility. No authorship demand by facility. User fee will be applied.
- Collaborative analysis: Besides research data acquisition (for which the same user fee will be applied as in Contract/service analysis), data analysis and interpretation will be provided. Archiving and public access will be provided. Intellectual property about the analysis and its results will be shared and authorship will be shared (please see our Rules on Intellectual Property and Publications).
The facility can also be engaged with Collaborative research, meaning that the facility is part of a grant application together with the collaborator(s) and all costs are already defined and given upfront of the analysis after grant approval.
Access for qualified personnel
Only for qualified personnel, it is possible to have access to LC-MS/MS instrumentation for chemical proteomics experiment performed directly by the user, for minimum 4 consecutive hours and with a mandatory initial start-up charge.
Visiting address
Karolinska Institutet
Department of Medical Biochemistry and Biophysics (MBB)
Biomedicum, quarter A9, floor 9
Solnavägen 9
Stockholm, Sweden
Contact
Massimiliano Gaetani
Head of FacilityPersonnel
Roman Zubarev, Ph.D. (Facility Director)
Massimiliano Gaetani, Ph.D. (Head of Facility)
Susanna Lundström, Ph.D. (Staff Senior Research Specialist in Proteomics)
Olga Lytovchenko, Ph.D. (Staff Research Engineer)
Zhaowei Meng (Staff LC-MS Engineer)
Xuepei Zhang, Ph.D. (Staff Research Specialist in Chemical Proteomics and HDX-MS)
Hassan Gharibi, Ph.D. (PostDoc, Data Analyst – Proteomics and Bioinformatics Expert)





