Small Ubiquitin Modifier (SUMO) signaling and muscle biology
The research group Small Ubiquitin Modifier (SUMO) signaling and muscle biology is lead by Stefano Gastaldello and located with the Department of Physiology and Pharmacology.
Research description
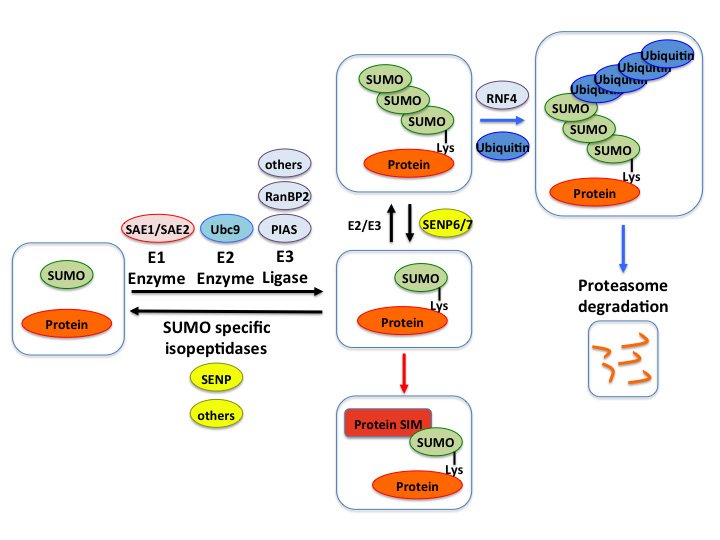
SUMO, the Small Ubiquitin-like Modifier is a 101-amino acid protein that can be covalently attached to lysine residues on target proteins via an enzymatic cascade reaction named SUMOylation.
SUMOylation is an important post-translational modification that regulates protein functions and contributes to several intracellular process including transcription, DNA repair, chromatin remodeling, signal transduction, etc. Particularly, various stress conditions are known to promote global changes in the SUMO signaling, both in cell cultures and at the organs level.
Moreover, defect on the SUMOylation machinery have been associated with severe diseases such as neurodegeneration, muscle atrophy, cancer and heart failure.
Our major contribution towards the muscle physiology field is to identify new skeletal muscle proteins targeted by SUMO in normal muscle activity, in muscle disorders and muscle-associated cancer as rhabdomyosarcoma.
Projects
i) The SUMO network in the skeletal muscle pathophysiology: from bench-to-bedside and beyond.
ii) The role of SUMO network during myogenesis in normal and in hyperglycemic/diabetes conditions.
iii) Alteration of the SUMO network in rhabdomyosarcoma cells - a new approach to study the proliferative and metastatic features of this tumor.
iv) Aging and SUMO network.

Group members
Stefano Gastaldello – Senior researcher, Group leader, Associate professor-Docent
Xiuxiu Liu – Master student, Binzhou Medical University, Yantai, China
Yizi Zhou - Master student, Binzhou Medical University, Yantai, China
Former members
Gabriel Heras Arribas, PhD 2020
Arvind Venkat Namuduri, PhD 2018
Silvia Codenotti, Post Doc, Department of Molecular and Translational Medicine (DMMT) University of Brescia, Italy
Selected publications
High glucose-induced oxidative stress accelerates myogenesis by altering SUMO reactions.
Liu X, Heras G, Lauschke VM, Mi J, Tian G, Gastaldello S
Exp Cell Res 2020 Oct;395(2):112234
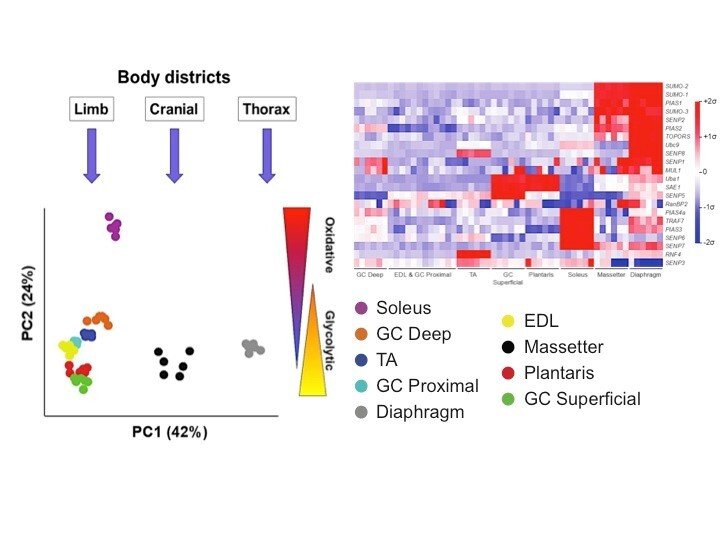
Expression of SUMO enzymes is fiber type dependent in skeletal muscles and is dysregulated in muscle disuse.
Namuduri AV, Heras G, Lauschke VM, Vitadello M, Traini L, Cacciani N, et al
FASEB J 2020 02;34(2):2269-2286
Muscle RING-finger protein-1 (MuRF1) functions and cellular localization are regulated by SUMO1 post-translational modification.
Heras G, Namuduri AV, Traini L, Shevchenko G, Falk A, Bergström Lind S, et al
J Mol Cell Biol 2019 05;11(5):356-370
Sarcolemmal loss of active nNOS (Nos1) is an oxidative stress-dependent, early event driving disuse atrophy.
Lechado I Terradas A, Vitadello M, Traini L, Namuduri AV, Gastaldello S, Gorza L
J Pathol 2018 12;246(4):433-446
A Proteomic Approach to Identify Alterations in the Small Ubiquitin-like Modifier (SUMO) Network during Controlled Mechanical Ventilation in Rat Diaphragm Muscle.
Namuduri AV, Heras G, Mi J, Cacciani N, Hörnaeus K, Konzer A, et al
Mol Cell Proteomics 2017 06;16(6):1081-1097
The chaperone co-inducer BGP-15 alleviates ventilation-induced diaphragm dysfunction.
Salah H, Li M, Cacciani N, Gastaldello S, Ogilvie H, Akkad H, et al
Sci Transl Med 2016 08;8(350):350ra103
Mechano-signalling pathways in an experimental intensive critical illness myopathy model.
Corpeno Kalamgi R, Salah H, Gastaldello S, Martinez-Redondo V, Ruas JL, Fury W, et al
J. Physiol. (Lond.) 2016 08;594(15):4371-88
Contact
Stefano Gastaldello
SUMO signalling and muscle biology group.
Associate Professor - Docent in Physiology. Department of Physiology and Pharmacology, Karolinska Institutet, Stockholm.
Visiting Professor at Faculty of Medicine and Pharmacy, Binzhou Medical University, Yantai, China.
Solnavägen 9, Quarter C5, 171 65 Stockholm, Sweden. Tel: + 46 (0) 8 524 87990. Skype: stefano.gastaldello.
