Embryonal, foetal and brain development - Juha Kere
Our knowledge of the very first few days of human life is still poor. We address questions such as: How does the embryo get started? How is the oocyte changed after fertilisation to gradually become mother of all different cell types in the body? Which genes regulate these earliest molecular mechanisms and how? What can we learn of stem cells and cellular reprogramming?
At the other end of foetal development, we study preeclampsia, a severe complication of pregnancies threatening both the mother and the baby. We look for diagnostic methods and therapy to better manage and prevent this serious disorder.
Early embryonic development and stem cells
Human embryo development starts by fertilisation of the egg cell (oocyte). The fertilised oocyte (zygote) will start to divide once a day, so that on day 2 the embryo has reached the 4-cell stage and on day 3, the 8-cell stage. The earliest events include the rapid but transient expression of the DUX4 gene in zygotes; the rapid degradation of a large number of oocyte-specific transcripts; and the activation of the embryo’s first own genes at the 4 and 8-cell stages.
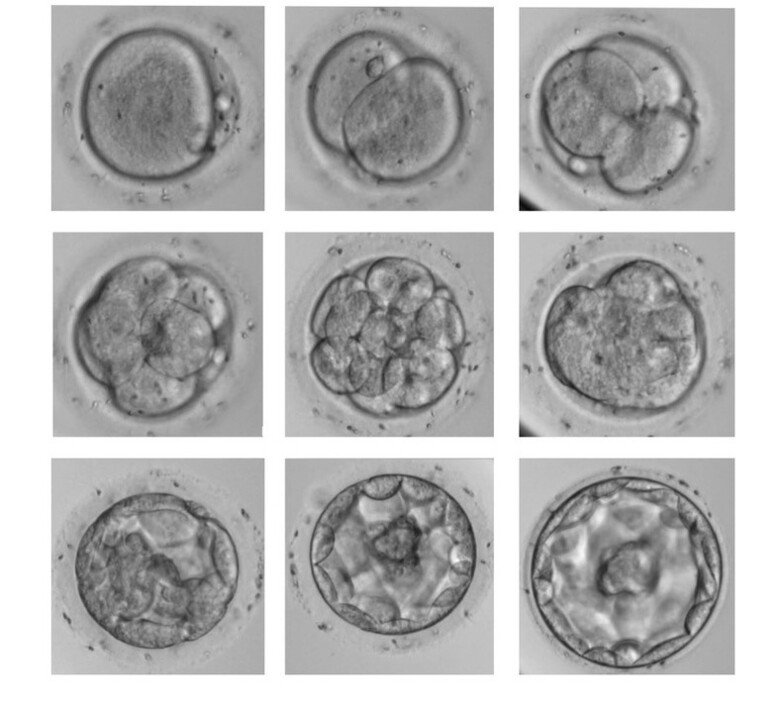
Recently, we have focused on the role of the DUX4 gene in regulating these processes. Our results have revealed multiple roles for DUX4, including a major effect in making the embryo genes accessible to regulatory proteins (chromatin opening) and activating a large number of enhancers, regulators of gene expression.
Preeclampsia, a severe complication of pregnancy
Preeclampsia is a serious pregnancy complication that starts during the last trimester of pregnancy and threatens the wellbeing of both the mother and the foetus. It is worldwide a leading cause of prematurity or even death during pregnancy. The aim of our project is to develop blood sample based sensitive methods to predict and diagnose threatening preeclampsia weeks before symptoms start. Our results show that an essential mechanism of preeclampsia involves a compromised genetic ability of the foetus to protect itself against maternal attack against the foreign body of placental cells. Our results have also suggested a therapy based on a well-known and safe medication.
Developmental dyslexia, a specific disorder of human brain development
Developmental dyslexia (DD), or specific reading disability, is an unexpected difficulty in learning to read despite normal intelligence and senses, and normal school teaching and social environment. DD is the most common learning disability, affecting between 5-10% of school-age children. Epidemiological studies have shown that DD has a strong genetic background. The aim of our project is to continue our groundbreaking research on the biological background of DD. After many gene discoveries, we now study gene functions in cell models. A key mechanism in DD involves signalling by neuronal cilia, antenna-like organelles that guide neuronal development.
Group members
Juha Kere
Group leaderTiina Skoog
Research EngineerHong Jiao
Affiliated to ResearchMasahito Yoshihara
Affiliated to ResearchNina Boskovic
Phd StudentGolbarg Vesterlund
Phd StudentSelected Publications
NET-CAGE characterizes the dynamics and topology of human transcribed cis-regulatory elements.
Hirabayashi S, Bhagat S, Matsuki Y, Takegami Y, Uehata T, Kanemaru A, Itoh M, Shirakawa K, Takaori-Kondo A, Takeuchi O, Carninci P, Katayama S, Hayashizaki Y, Kere J, Kawaji H, Murakawa Y. Nature Genet 51:1369-1379, 2019
Pleomorphic Adenoma Gene 1 is needed for timely zygotic genome activation and early embryo development.
Madissoon E, Damdimopoulos A, Katayama S, Krjutškov K, Einarsdottir E, Mamia K, De Groef B, Hovatta O, Kere J*, Damdimopoulou P*. Sci Rep 9:8411, 2019
Human pluripotent reprogramming with CRISPR dCas9 activators.
Weltner J, Balboa D, Katayama S, Bespalov M, Krjutškov K, Jouhilahti E-M, Trokovic R, Kere J*, Otonkoski T*. Nature Commun 9:2643, 2018
The human PRD-like homeobox gene LEUTX has a central role in embryo genome activation.
Jouhilahti E-M, Madissoon E, Vesterlund L, Töhönen V, Krjutshkov K, Plaza Reyes A, Petropoulos S, Månsson R, Linnarsson S, Bürglin T, Lanner F, Hovatta O, Katayama S, Kere J. Development 143:3459-3469, 2016
Research Networks
- EU Consortia: MAARS, SARM, NANOSOL, BIOMAP, MATER
- Other international consortia (coordinator country): FANTOM6 (Japan), FINNPEC (Finland)
Prizes/Awards
- Juha Kere: The Royal Society Wolfson Research Merit Award (UK)
